The R@CMon team has been working closely with the GVL (Genomics Virtual Laboratory) team to bring the GVL to the Monash node of the NeCTAR Research Cloud. It’s now here!
The GVL provides an accessible infrastructure for connecting genome researchers with their datasets using analysis tools that are executed on dynamically provisioned computational resources – Amazon AWS EC2 and the NeCTAR Research Cloud. The GVL Launcher allows users to launch CloudBioLinux, CloudMan and Galaxy computational platforms on a number of cloud computing infrastructures.
Initially, the GVL was only available on the Melbourne node. Persistent volume storage is one of the main requirements of the GVL. With the initial volume service now available on the Monash node, users can now launch the GVL at the Monash node. By choosing the Monash availability zone, the GVL head node and all subsequent compute nodes will be launched at monash-01.
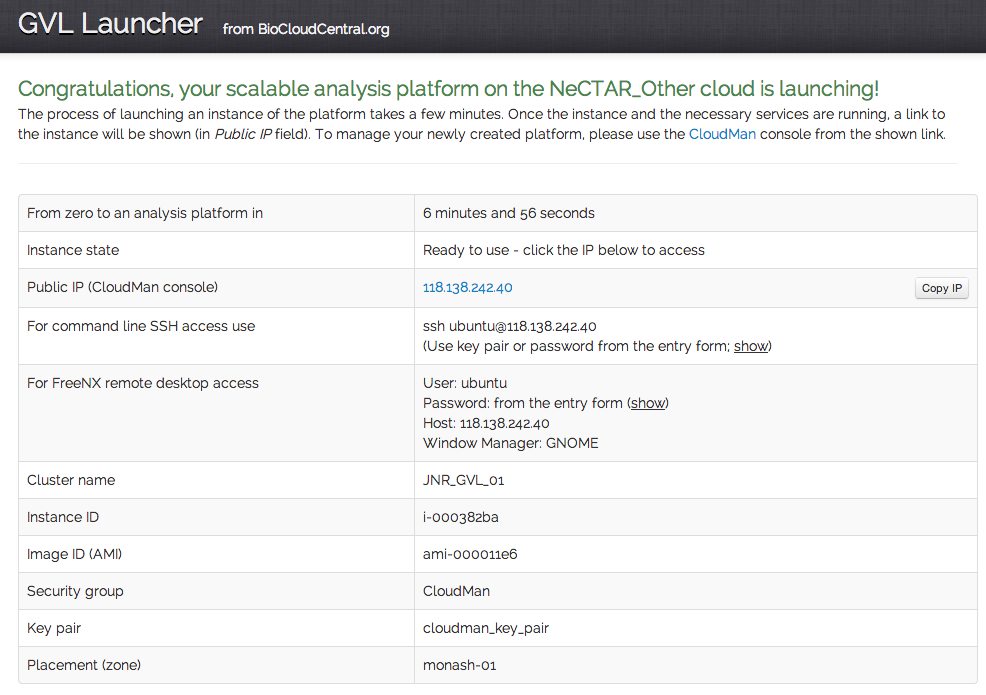
The GVL team makes the latest stable GVL image available to the users. And now, we’re ready to launch our first GVL platform in the Monash node. This will launch the head node for the platform. It will be accessible via CloudMan web interface once it has been booted up and all necessary services have been initialised. In the next parts of this blog series, the various GVL use cases will be covered, including – using a dynamically scaled GVL platform.